Homolog information
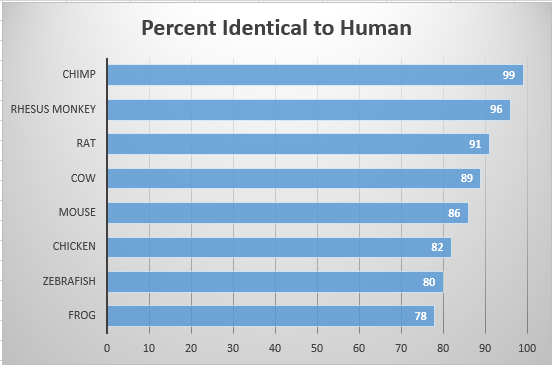
Identifying homologs can be a helpful genetic tool for finding good model organisms. In order to determine the homologs of the GABRB3 protein, the National Center for Biotechnology Information's program, BLAST, was used. The NCBI Homologene database was also utilized. Homologene constructs homology groups from gene sets of different species. Separate sequences are compared, and similarities between organisms are found. Eight homologs were identified for this analysis. As more information is gathered, more homologs may be identified and added to the list below.
As indicated by the graph, the GABRB3 gene is well conserved across many different species. As expected, mammals and vertebrates are most similar to the human model, and the fly and roundworm are least similar, although still well-conserved. Many of these species would make good model organisms for experiments involving GABRB3. It is important to conduct experiments with organisms that reflect the target model well (in this case, humans), so that any advanced trials will yield similar results.
|
Human (Homo sapien)
(GABA) A receptor, beta 3 Accession Number: NM_001191321.2 GI Number: 522838290 FASTA |
Chimp (Pan Troglodyte)
(GABA) A receptor, beta 3 Accession Number: XM_003314572.1 GI Number: 332843340 FASTA |
|
Mouse (Mus musculus)
(GABA) A receptor, beta 3 Accession Number: NM_001038701.2 GI Number: 551410328 FASTA |
Zebrafish (Danio rerio)
(GABA) A receptor subunit, beta 2 -like Accession Number: XM_005174449.1 GI Number: 528523332 FASTA |
|
Rhesus Monkey (Macaca Mulatta)
(GABA) A receptor, beta 3 Accession Number: NM_00126702.1 GI Number: 386781492 FASTA |
|
Rat (Rattus Norvegicus)
(GABA) A receptor, beta 3 Accession Number: NM_017065.1 GI Number: 8393389 FASTA |
Chicken (Gallus Gallus)
(GABA) A receptor, beta 3 Accession Number: NM_205346.3 GI Number: 482677673 FASTA |
|
Frog (Xenopus)
(GABA) A receptor, subunit beta-4-like Accession Number: XM_004916771.1 GI Number: 512858729 FASTA |
Phylogeny
Phylogenetics revolves around studying the relationship between species. The concept of a phlylogeny tree was first introduced by Charles Darwin, and has since been developed into a useful evolutionary tool. In order to reconstruct evolutionary history, we need to identify homologous characters that are shared between species.
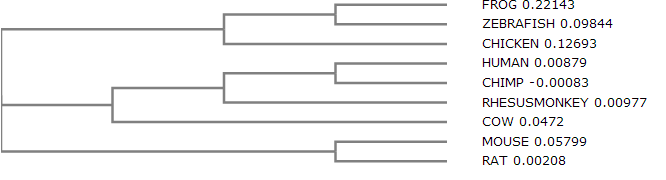
In order to build this phylogenetic tree, the program Clustal Omega was used. Clustal Omega is a multiple sequence alignment program that uses seeded guide trees and HMM profile-profile techniques to generate alignments.
In order to build this phylogenetic tree, the program Clustal Omega was used. Clustal Omega is a multiple sequence alignment program that uses seeded guide trees and HMM profile-profile techniques to generate alignments.
As the phylogram indicates, the human GABRB3 DNA is most closely related to that of the chimp and rhesus monkey. These results are supported by the Percent Identity graph data comparing different organisms to the human DNA. The model organism I was most interested in for my specific aims was the mouse, because of its close relation to the human.
References:
1. Delsuc F, Brinkmann H, Philippe H. Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet. 2005 May;6(5):361-75. doi: 10.1038/nrg1603.
2. Clustal Omega: http://www.ebi.ac.uk/Tools/msa/clustalo/
3. BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi
1. Delsuc F, Brinkmann H, Philippe H. Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet. 2005 May;6(5):361-75. doi: 10.1038/nrg1603.
2. Clustal Omega: http://www.ebi.ac.uk/Tools/msa/clustalo/
3. BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi